Слайд 2Two methods are widely used to determine the concentration of
DNA in solution. The most simple and accurate is the
spectrophotometric method, but it has a relatively low sensitivity. If the total content of nucleic acids is small, the concentration of DNA can be determined from the intensity of their fluorescence in UV light after staining with ethidium bromide
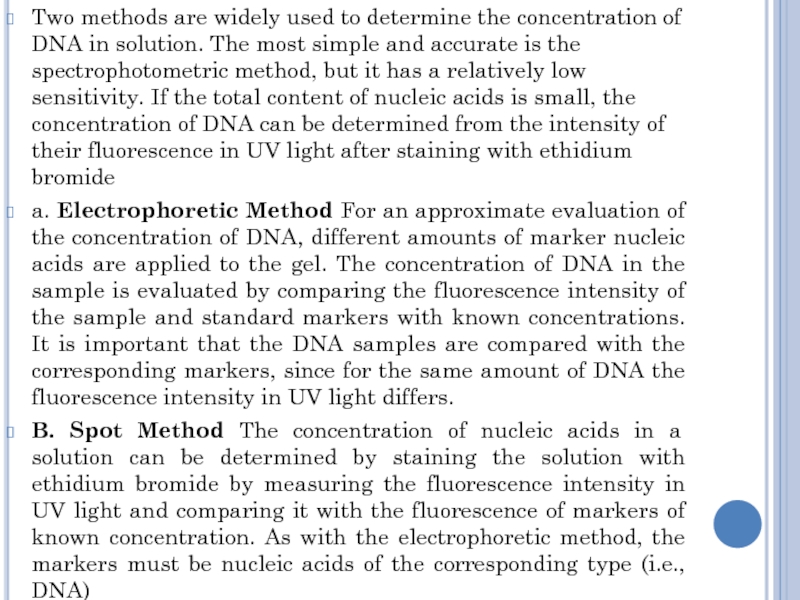
a. Electrophoretic Method For an approximate evaluation of the concentration of DNA, different amounts of marker nucleic acids are applied to the gel. The concentration of DNA in the sample is evaluated by comparing the fluorescence intensity of the sample and standard markers with known concentrations. It is important that the DNA samples are compared with the corresponding markers, since for the same amount of DNA the fluorescence intensity in UV light differs.
B. Spot Method The concentration of nucleic acids in a solution can be determined by staining the solution with ethidium bromide by measuring the fluorescence intensity in UV light and comparing it with the fluorescence of markers of known concentration. As with the electrophoretic method, the markers must be nucleic acids of the corresponding type (i.e., DNA)

Слайд 3Nucleic acids
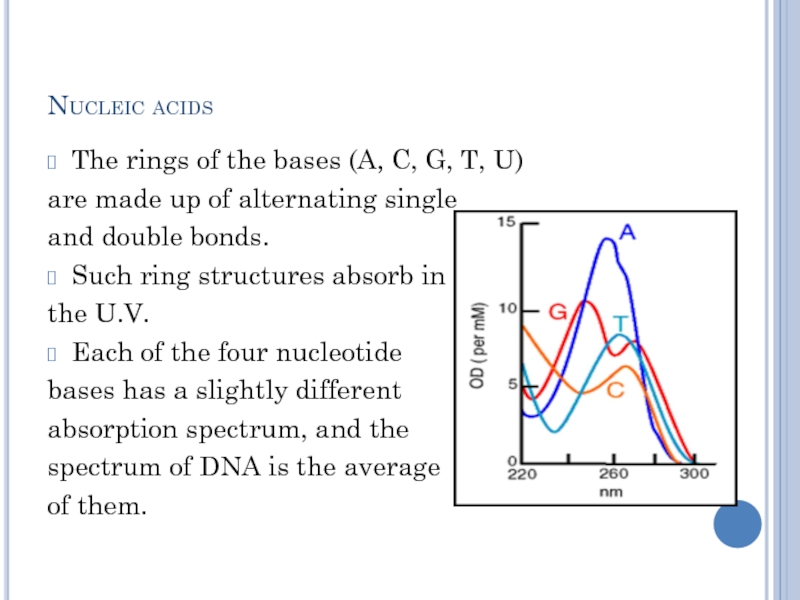
The rings of the bases (A, C, G, T,
U)
are made up of alternating single
and double bonds.
Such ring structures absorb in
the U.V.
Each of the four nucleotide
bases has a slightly different
absorption spectrum, and the
spectrum of DNA is the average
of them.
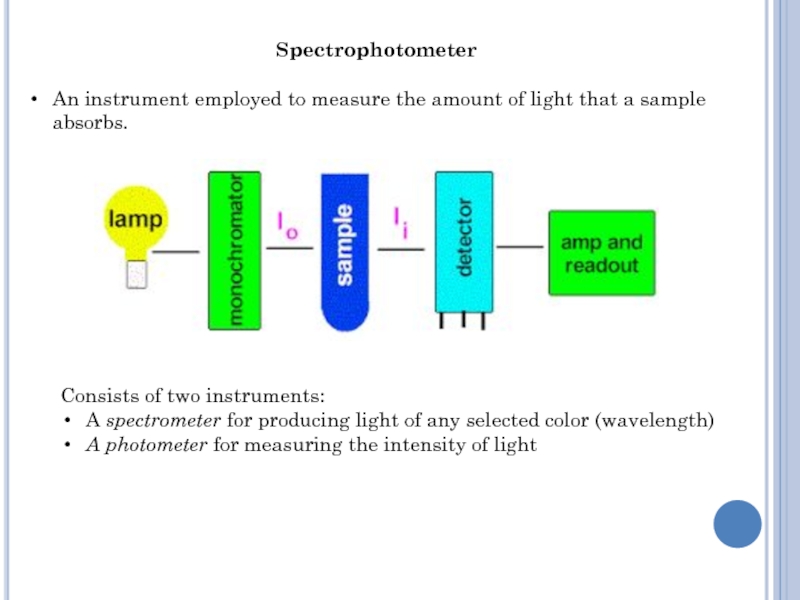
Слайд 4Spectrophotometer
An instrument employed to measure the amount of light that
a sample absorbs.
Consists of two instruments:
A spectrometer for producing
light of any selected color (wavelength)
A photometer for measuring the intensity of light
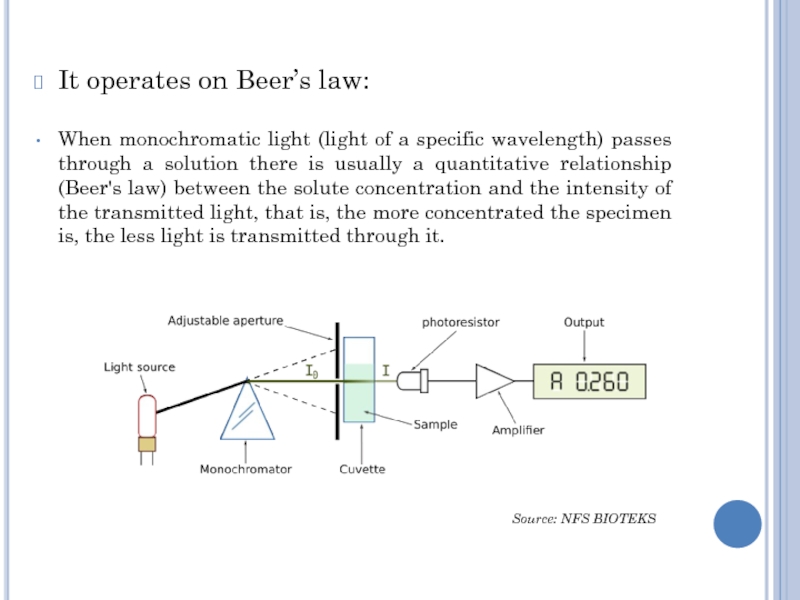
Слайд 5It operates on Beer’s law:
When monochromatic light (light of a
specific wavelength) passes through a solution there is usually a
quantitative relationship (Beer's law) between the solute concentration and the intensity of the transmitted light, that is, the more concentrated the specimen is, the less light is transmitted through it.
Source: NFS BIOTEKS
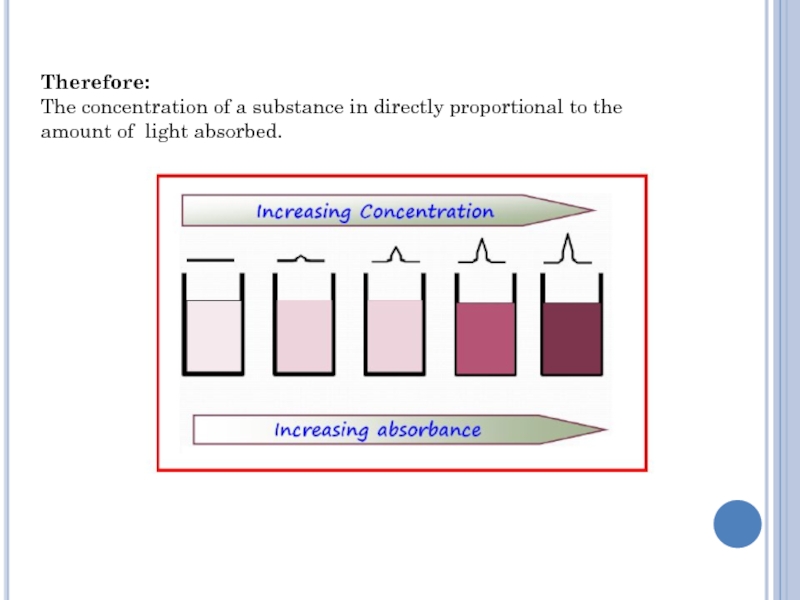
Слайд 6Therefore:
The concentration of a substance in directly proportional to
the
amount of light absorbed.
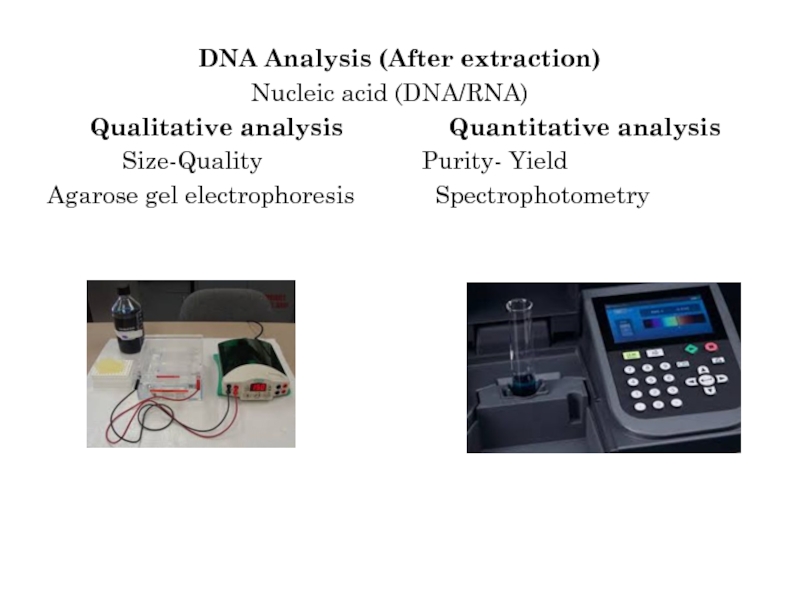
Слайд 7DNA Analysis (After extraction)
Nucleic acid
(DNA/RNA)
Qualitative analysis Quantitative analysis
Size-Quality Purity- Yield
Agarose gel electrophoresis Spectrophotometry
Слайд 8
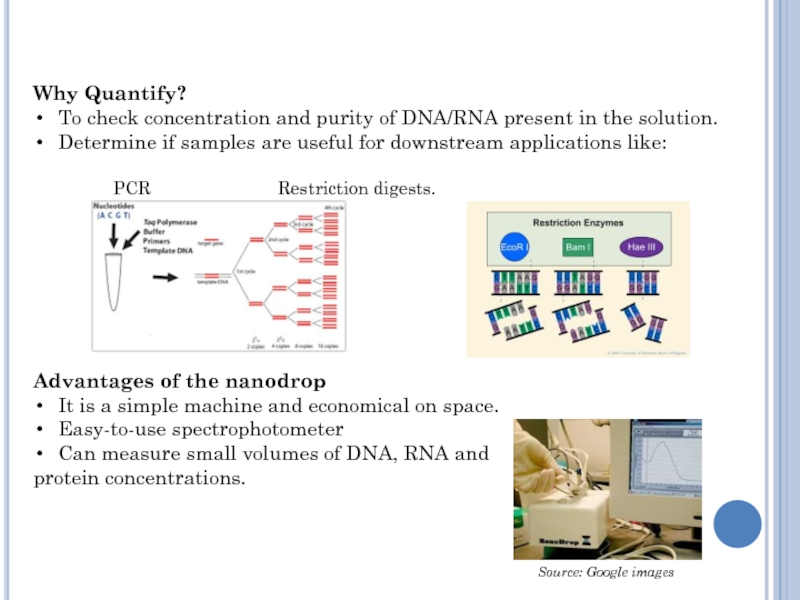
Why Quantify?
To check concentration and purity of DNA/RNA present in
the solution.
Determine if samples are useful for downstream applications like:
PCR Restriction digests.
Advantages of the nanodrop
It is a simple machine and economical on space.
Easy-to-use spectrophotometer
Can measure small volumes of DNA, RNA and
protein concentrations.
Source: Google images
Слайд 9Spectrophotometric measures
Measures DNA, RNA (A260) and Proteins (A280) concentrations and
sample purity (260:280).
Absorbance at 260 nm
Nucleic acids absorb UV light
at 260 nm due to the aromatic base moieties
within their structure. Purines (thymine, cytosine and uracil) and pyrimidines (adenine and guanine) both have peak absorbances at 260 nm, thus making it
the standard for quantitating nucleic acid samples.
Absorbance at 280 nm
The 280 nm absorbance is measured where proteins and phenolic compounds have a strong absorbance. Similarly, the aromaticity of phenol groups of organic compounds absorbs strongly near 280 nm.
Absorbance at 230 nm
Many organic compounds have strong absorbances at around 225 nm. In addition to phenol, TRIzol, and chaotropic salts, the peptide bonds in proteins absorb light between 200 and 230 nm.
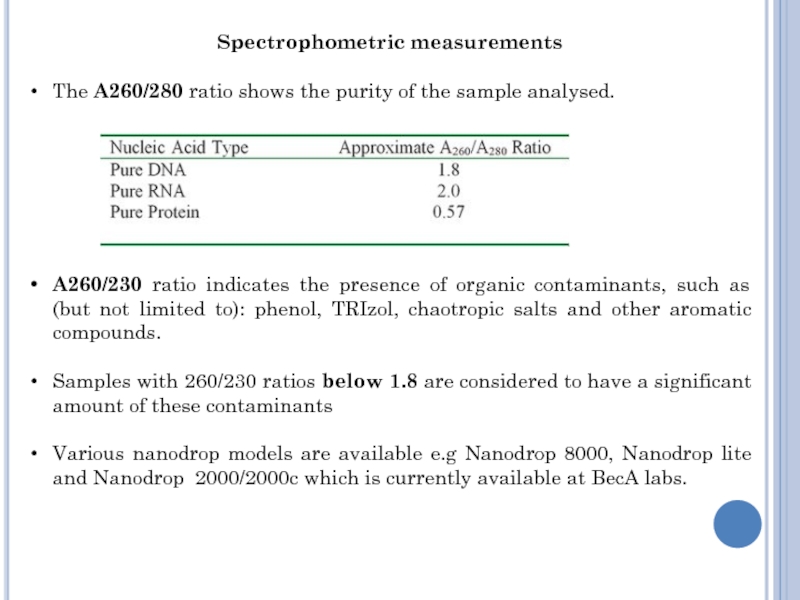
Слайд 10Spectrophometric measurements
The A260/280 ratio shows the purity of the sample
analysed.
A260/230 ratio indicates the presence of organic contaminants, such as
(but not limited to): phenol, TRIzol, chaotropic salts and other aromatic compounds.
Samples with 260/230 ratios below 1.8 are considered to have a significant amount of these contaminants
Various nanodrop models are available e.g Nanodrop 8000, Nanodrop lite and Nanodrop 2000/2000c which is currently available at BecA labs.
Слайд 11Note that:
Because of the low concentrations, sometimes it is difficult
to assess the purity of the samples (esp. RNA) by
analyzing the A260/280 and A260/230 ratios.
Quantification cannot be assessed by the NanoDrop because they are outside the lowest concentrations the NanoDrop is designed to measure.
A more sensitive method such as an Agilent Biolanalyzer or Qubit analysis is recommended.
Слайд 12Electrophoresis is a method whereby charged molecules in solution, chiefly
proteins and nucleic acids, migrate in response to an electrical
field.
Their rate of migration through the electrical field, depends on the strength of the field, on the net charge, size, and shape of the molecules, and also on the ionic strength, viscosity, and temperature of the medium in which the molecules are moving.
As an analytical tool, electrophoresis is simple, rapid and highly sensitive.
It can be used analytically to study the properties of a single charged species or mixtures of molecules. It can also be used preparatively as a separating technique
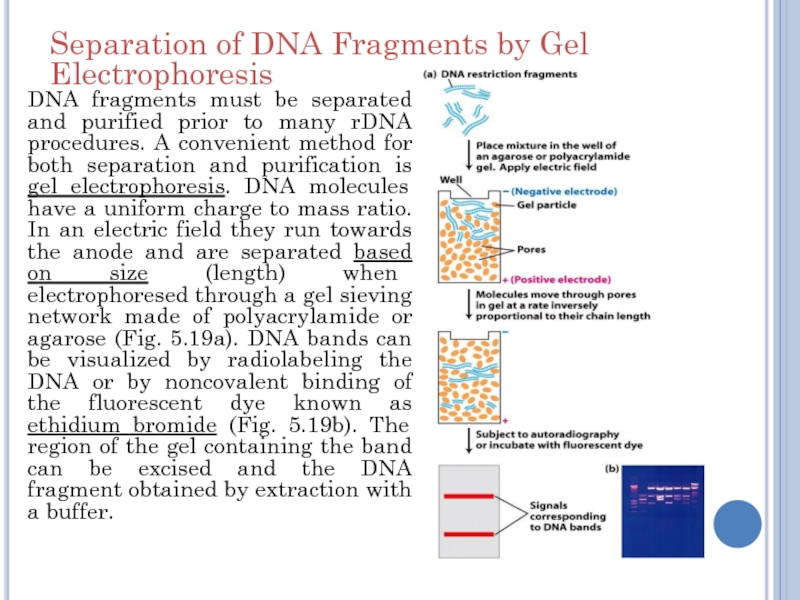
Слайд 13DNA fragments must be separated and purified prior to many
rDNA procedures. A convenient method for both separation and purification
is gel electrophoresis. DNA molecules have a uniform charge to mass ratio. In an electric field they run towards the anode and are separated based on size (length) when electrophoresed through a gel sieving network made of polyacrylamide or agarose (Fig. 5.19a). DNA bands can be visualized by radiolabeling the DNA or by noncovalent binding of the fluorescent dye known as ethidium bromide (Fig. 5.19b). The region of the gel containing the band can be excised and the DNA fragment obtained by extraction with a buffer.
Separation of DNA Fragments by Gel Electrophoresis
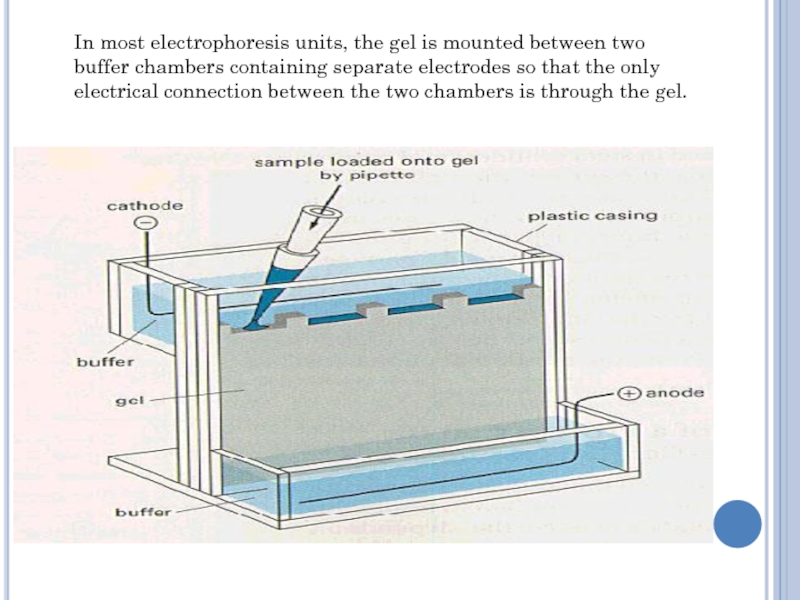
Слайд 14In most electrophoresis units, the gel is mounted between two
buffer chambers containing separate electrodes so that the only electrical
connection between the two chambers is through the gel.
Слайд 15Although agarose and polyacrylamide differ greatly in their physical and
chemical structures, they both make porous gels.
A porous gel
acts as a sieve by retarding or, in some cases, by completely obstructing the movement of macromolecules while allowing smaller molecules to migrate freely.
By preparing a gel with a restrictive pore size, the operator can take advantage of molecular size differences among proteins
Слайд 16Fluorescent dyes are multicyclic molecules that absorb and emit fluorescent
light at specific wavelengths.
Examples are fluorescein and rhodamine derivatives.
For sequencing applications, these molecules can be covalently attached to nucleotides.
Слайд 17In dye primer sequencing, the primer contains fluorescent dye–conjugated nucleotides,
labeling the sequencing ladder at the 5′ ends of the
chains.
In dye terminator sequencing, the fluorescent dye molecules are covalently attached to the dideoxynucleotides, labeling the sequencing ladder at the 3′ ends of the chains.
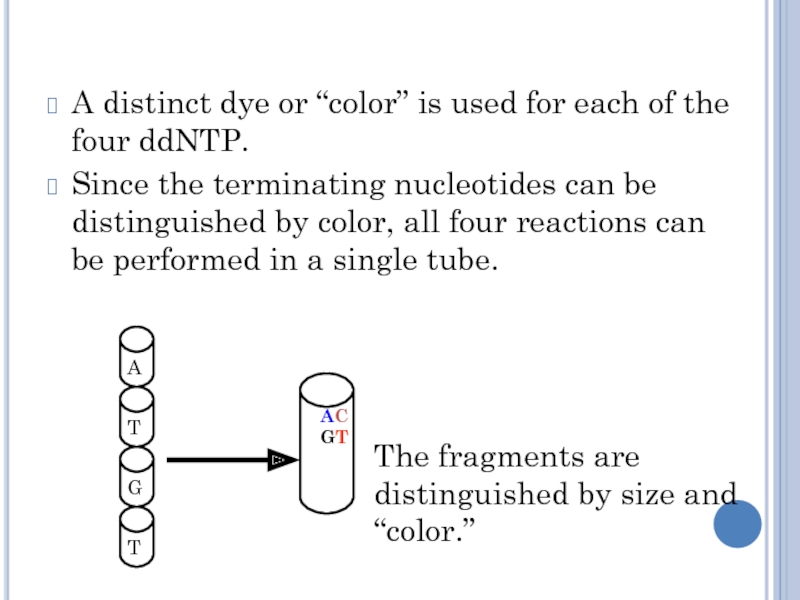
Слайд 18A distinct dye or “color” is used for each of
the four ddNTP.
Since the terminating nucleotides can be distinguished by
color, all four reactions can be performed in a single tube.
A
T
G
T
The fragments are distinguished by size and “color.”
Слайд 19
5′ AGTCTG
5′ AG(T/A)CTG
5′ AGACTG
T/T
T/A A/A
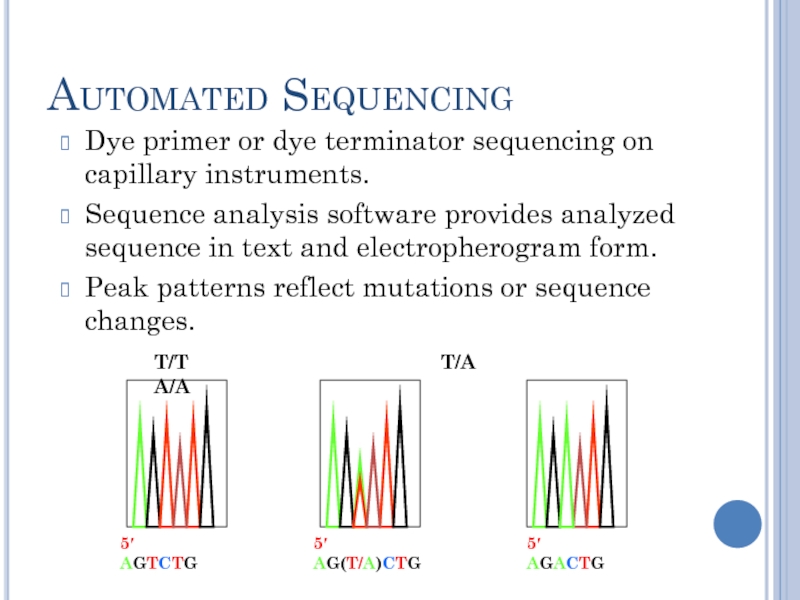
Automated Sequencing
Dye primer or dye terminator sequencing on capillary instruments.
Sequence analysis software provides analyzed sequence in text and electropherogram form.
Peak patterns reflect mutations or sequence changes.